1928 is "easy-to-use"
2021-06-21
When we interact with our users we get fantastic quotes on the ease of use of our service. Some say it is “idiot-proof”. We say it is user-friendly.
Today, hospitals often lack bioinformatic resources that are necessary to take advantage of available outbreak data, such as sequences published by public health authorities or even sequences from samples taken in their own clinics. The 1928 Diagnostics service fills this gap by offering bioinformatic analysis of bacteria, providing species identification, antibiotic resistance profiling, and genetic relatedness for transmission management- all in a matter of minutes.
In Slott Jensen et al.’s letter to the editor, published June 9th, 2021 in Infection Control & Hospital Epidemiology, the authors emphasize the usability of the 1928 platform. They state that
“Rapid typing of selected bacteria is essential for effective surveillance and early detection of outbreaks in hospitals. The simple procedure for uploading data and viewing results makes 1928 Diagnostics attractive to infection control units without bioinformatic expertise.”
You can read the entire letter here.
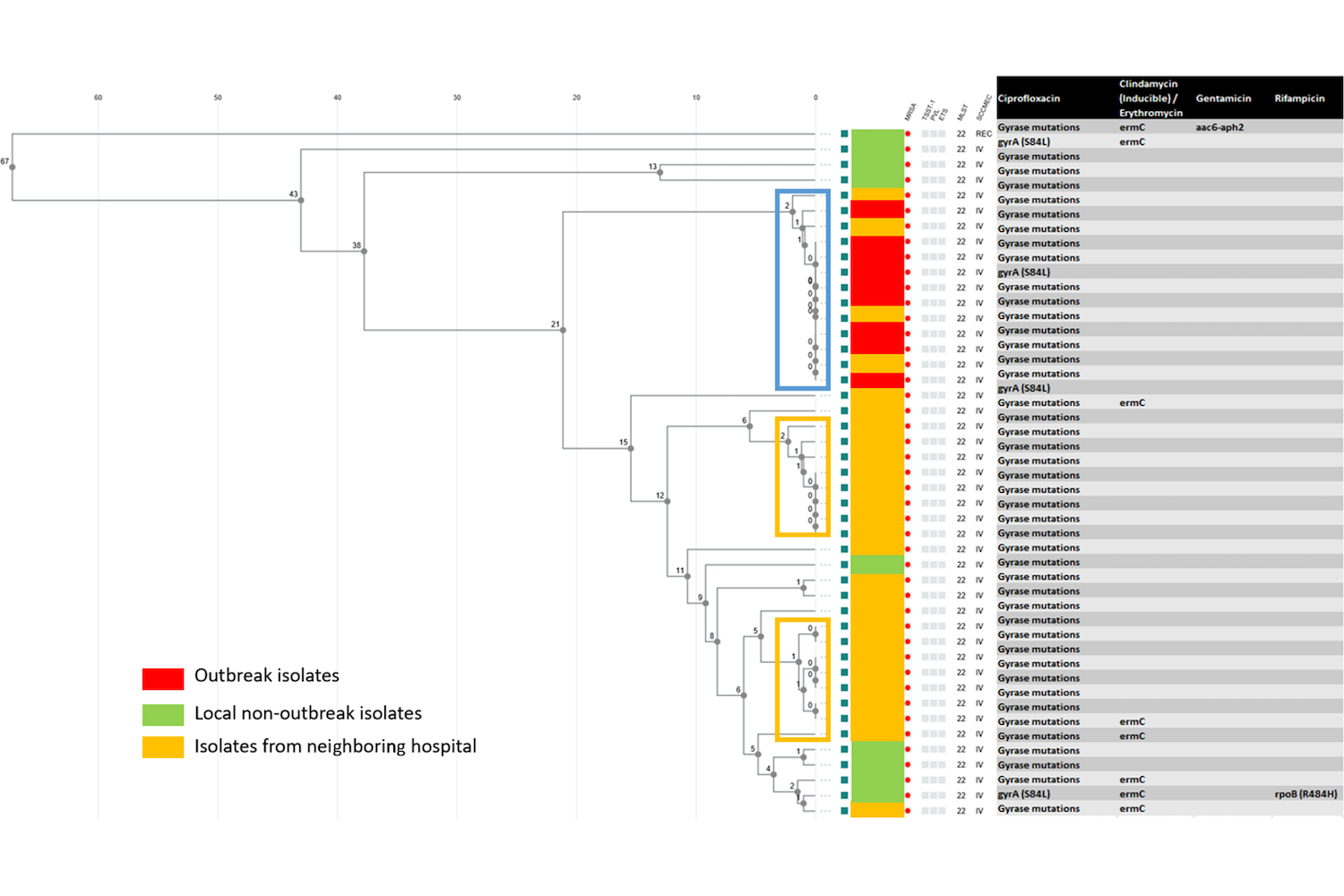
Slott Jensen, M., Chen, M., Detlefsen, M., Kudsk Klitgaard, J., Andersen, T., & Kemp, M. (2021). Whole-genome sequence analyses by a new easy-to-use software solution support the suspicion of a neonatal ward outbreak of methicillin-resistant Staphylococcus aureus (MRSA) and transmission between hospitals. Infection Control & Hospital Epidemiology, 1-3. doi:10.1017/ice.2021.123
Curious to try: